Welcome to CAMBer project website
We recommend you to visit the website of eCAMBer - our new software, which signifficantly optimizes and extends CAMBer. (paper submitted to Bioinformatics, 2013).
A large amount of genomic data is now publicly available, enabling very interesting comparative analysis. However, we do observe a lot of inconsistencies in the genome structure annotations among bacterial strains (John Dunbar et al. BMC Genomics, 12(1):125, 2011). This inconsistency is a frustrating impedance to effective comparative genomic analysis of bacterial strains in promising applications such as gaining insights into bacterial drug resistance.
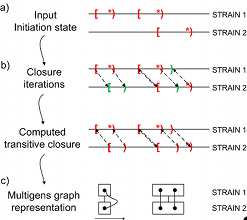
We developed CAMBer as an approach to support comparative analysis of multiple bacterial strains (Wozniak et al. BMC Genomics 2011, 12(Suppl 2):S6). CAMBer unifies annotations of closely related species by homology transfer. It produces what we called multigene families. Each multigene family reveals genes that are in one-to-one correspondence in the bacterial strains, thereby permitting their annotations to be integrated. In our paper we present more details and results of our method applied to three human pathogens: Escherichia coli, Mycobacterium tuberculosis and Staphylococcus aureus.
In order to visualize inconsistencies in genome annotations detected by CAMBer we have developed a visualization tool, CAMBerVis (Wozniak et al. Bioinformatics 2011). The software manages simultaneous visualization of multiple bacterial genomes, enabling visual analysis focused on inconsistencies in genome structure annotations.