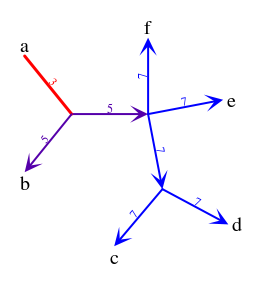
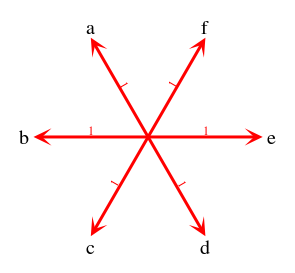
Robinson-Foulds - unrooted gene trees vs rooted species treesContents
General description
This page contains complementary material for the article [1]:
prototype software and results of experiments.
Program
Program rf in package rf.tgz have the
following options:
rf [h] [-g FILE|-] [-s FILE|-] [FILE1] [FILE2] ... FILE can be: - (minus) - stdin, filename or a string (if file of given name does not exists). Option -g defines gene trees, option -s species trees. Trees are defined in standard nested parenthesis notation. If no option is used rf assumes that the first tree is a gene tree, then a species tree, a gene tree, etc.
Output. Nodes are decorated with atributes:
Downloads
All files can be accessed from this directory: files.
Bibliography
[1] Pawel Gorecki and Oliver Eulenstein, A Robinson-Foulds measure to compare unrooted trees with rooted trees
Homepage
|